 PLATINUM is designed for calculation of hydrophobic properties of molecules and their match or mismatch in receptor–ligand complexes. These properties may help to analyze results of molecular docking.
PLATINUM is designed for calculation of hydrophobic properties of molecules and their match or mismatch in receptor–ligand complexes. These properties may help to analyze results of molecular docking.
Step 1: Upload files
Please select a ’ligand“ file (in one-by-one mode) — if you want just to assess their hydrophobic properties — and add a single receptor file to assess ligand/receptor interactions. (File size limit: 5 Mb.)
Please note: PLATINUM requires that all molecules are protonated!
What is PLATINUM?
Calculate and visualize molecular hydrophobic/hydrophilic properties using the concept of “Molecular Hydrophobicity Potential” (MHP). Hydrophobic and stacking interactions in biomolecular complexes can be estimated and used
How to use PLATINUM?
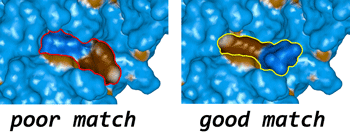
- Complementarity of hydrophobic/hydrophilic properties in biomolecular complexes:
- to view molecular hydrophobic properties upload ligand(s);
- to view hydrophobic complementarity add a receptor;
- on Step 2 select a reference ligand (optionally).
- 2D hydrophobic maps for lipid bilayers and α-helices:
- upload a bilayer or an α-helix as a receptor (currently only.GRO files are allowed);
- add ligand(s) to view the contact area on the map (optionally).
For details refer to the manual and go through the tutorial; don’t forget files for the tutorial.
System requirements
Some components on the site (viz. massive files upload) require modern HTML5-capable web-browser with javascript enabled.
